Genomics:
Gene, Genome, and Gene Expression Research
Mitochondrial Genomes in ME/CFS
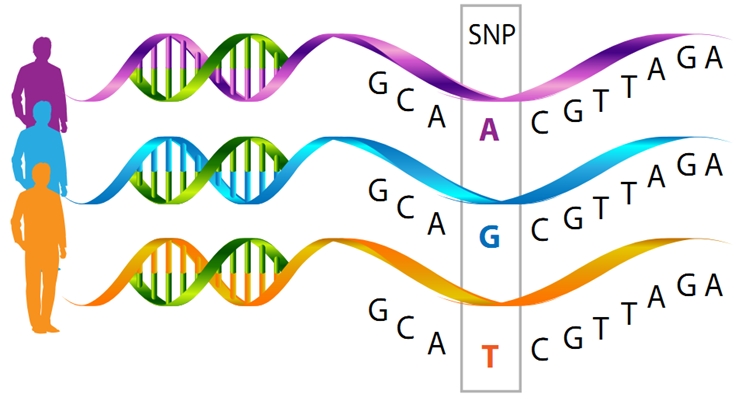
Mitochondria are made up of molecules encoded by the nuclear genome--DNA located in the nucleus--as well as the mitochondrial genome—a small amount of DNA present within each organelle. Defects in mitochondrial DNA lead to devastating genetic diseases, with such symptoms as brain abnormalities, severe fatigue, blindness or defective heart function—and can be fatal. The mitochondrial genome of healthy humans also exhibits some natural variation—a single component of the mitochondrial DNA sometimes differs between one human and another—this is known as a SNP (single nucleotide polymorphism, "snip"). Often more than one SNP differs between one population of humans and another—for example, mitochondrial genomes whose origin can be traced to France differ in a number of SNPs from those in people in Central Asia. These different types of mitochondrial genomes, based on a specific set of SNPs, are referred to as haplogroups. Even people whose mitochondrial DNA belongs to the same haplogroup can differ among one another because of some variation in additional SNPs. Some mitochondrial SNPs have been associated with various characteristics, such as adaptation to cold weather or high altitude environments and have been implicated in susceptibility to diabetes and various inflammatory diseases.
A further complexity of mitochondrial genetics is that there are many individual mitochondria within the same cell, and thus many copies of mitochondrial DNA in each cell. Sometimes new mutations arise so that some of the copies of DNA within the same cell, and therefore within the same person, differ from one another. This situation is called “heteroplasmy”. As cells grow and multiply, by chance there can be uneven distribution of normal vs. abnormal DNA to different cells. If mitochondrial DNA with a harmful mutation becomes the predominant type in a particular tissue, serious symptoms will emerge.
 With funding by the Chronic Fatigue Initiative, the lab groups of Zhenglong Gu, Alon Keinan, and Maureen Hanson collaborated to sequence and analyze mitochondrial DNAs in 193 patients and 196 controls. Patients were selected and samples collected by the ME/CFS expert physicians working with the Chronic Fatigue Initiative.
With funding by the Chronic Fatigue Initiative, the lab groups of Zhenglong Gu, Alon Keinan, and Maureen Hanson collaborated to sequence and analyze mitochondrial DNAs in 193 patients and 196 controls. Patients were selected and samples collected by the ME/CFS expert physicians working with the Chronic Fatigue Initiative.
As described in the publication listed below, none of the patients had pathogenic mitochondrial DNA mutations. There was no difference in the extent of heteroplasmy between ME/CFS patients and controls. No increased susceptibility to ME/CFS among individuals carrying particular haplogroups or SNPs within a haplogroup was detected.
Statistically significant correlations of particular SNPs could be made with certain symptoms and/or their severity. However, the sample size of 393 subjects is relatively small, and these findings require further validation with additional subjects.
Additional information:
Billing-Ross, P., Germain A. , Ye K., Keinan A., Gu Z, and M. R. Hanson 2016.
Mitochondrial DNA variants correlate with symptoms in myalgic encephalomyelitis/chronic fatigue syndrome.
J. Translational Medicine. 14:19.
 Center for Enervating NeuroImmune Disease
Center for Enervating NeuroImmune Disease